BLAST Highlights
NCBI has recently designed the BLAST interface to improve usability.
New features include:
- A list of search results performed within the past 36 hours
- Search strategies that can be saved to MyNCBI
- Reorganized home-page links to BLAST forms
- Redesigned BLAST forms with usability enhancements
- A documentation catalog
Common Interface Elements
All pages in the new design have the same page header. The page header always has the standard MyNCBI login box at the top right, and a link to the NCBI home page at the top left.
The footer of every page has links to NCBI's copyright, disclaimer, privacy, accessibility pages; a contact page; and links to the home pages of NCBI and its parent organizations.
The header of each page features four tabs:

- Home Tab: Links to BLAST home page
- Recent Results Tab: Links to searches that you have performed in the past 36 hours
- Saved Strategies Tab: Filled-in BLAST input forms that you have saved to MyNCBI
- Help Tab: A catalog of BLAST documentation
Home Tab
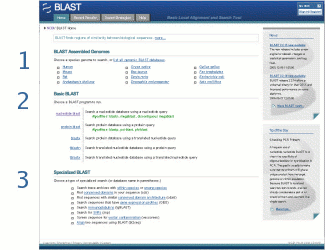
The Home tab always leads back to the BLAST home page. The narrow right column of the page contains News and Tip of the Day features that change periodically.
The wide left column of the page contains three sections:
- BLAST Assembled Genomes contains links to genomic BLAST pages for common organisms, and a link to a complete list of available organism genome BLAST pages
- Basic BLAST contains links to BLAST forms for the traditional set of databases (e.g., nr, est, etc.). Choose the link for the search type you want. For example, choose "nucleotide blast" to search a nucleotide database using a nucleotide query. Descriptions of the search types appear to the right of the links. See BLAST Forms below for a description of key features of these forms.
- Specialized BLAST contains links to special-purpose BLAST databases and tools such as trace archives and IgBLAST.
Recent Results Tab
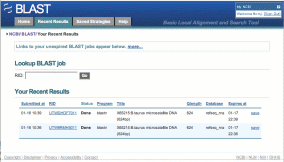
Recent Results provides links to searches performed within the past 36 hours. If you are signed in to MyNCBI, links to these searches will be available anywhere else you sign in. If you are not signed in, the links are only for those jobs run in that browser session, and disappear when the browser is closed. MyNCBI automatically saves all unexpired links when you sign in. Sign in or register for a free account by using the MyNCBI box at the top right of the page.
You may also look up any BLAST job by RID from this tab. BLAST RIDs are now 11 characters long (instead of 37), making them easier to jot down, read over the phone, or manually enter into programs.
Saved Strategies Tab
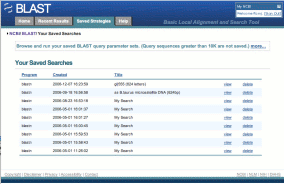
Saved Strategies allows you to save the parameters for a BLAST search so you can run the same job again in the future. This feature requires you to be signed in to MyNCBI.
Help Tab
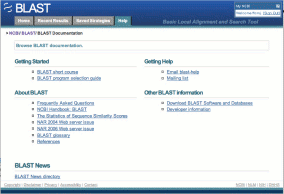
Help provides a catalog of links to documentation and publications about BLAST.
BLAST Forms
The links in the Basic BLAST section of the Home tab lead to BLAST forms that share a common design. Each form has four sections (examples are from nucleotide BLAST):
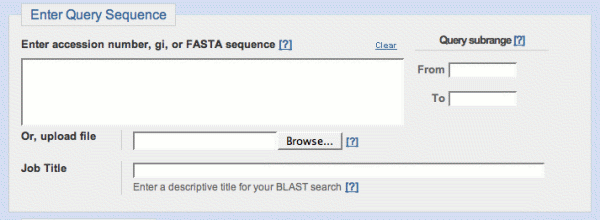
- Enter Query Sequence provides a place to input or
upload your query sequence, and optionally select a query subrange. You
may also supply a title for the job, which will appear at the top of
the corresponding BLAST report. A default title is automatically
created for each search.
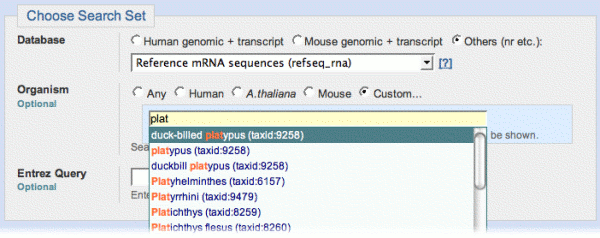
- Choose Search Set is where you select a database and optionally limit your search by an organism or Entrez query. The default database "Human genomic + transcript", implicitly limits the search to Human. If you select a database that is not species-specific (e.g., refseq_rna), an "Organism" control appears that lets you limit your search by organism. The "Custom" organism radio button brings up an auto-complete text box where you can type the (scientific or common) name or taxid of an organism to use as a limit.
- Program selection allows you to optimize your search for different scenarios (e.g., intra- vs. inter-species searches). The choices correspond to megablast, discontiguous megablast, and blastn for nucleotide; and blastp, PSI-BLAST and PHI-BLAST for protein.
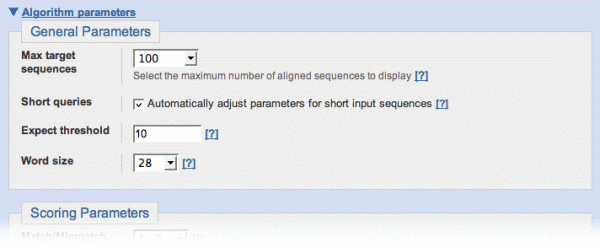
- Algorithm parameters is a link to a page section that lets you change the parameters of the selected BLAST algorithm. Most users don't need to see or change these parameters, so this section is usually "closed". Click the link to "open" the section.

Help Us Improve BLAST
Please send feedback on the BLAST redesign to blast-help@ncbi.nlm.nih.gov.
More Information
For more about BLAST, see the BLAST documentation directory.